Ashkaan’s paper was accepted in AEM!
https://www.ncbi.nlm.nih.gov/pubmed/28411219
ABSTRACT
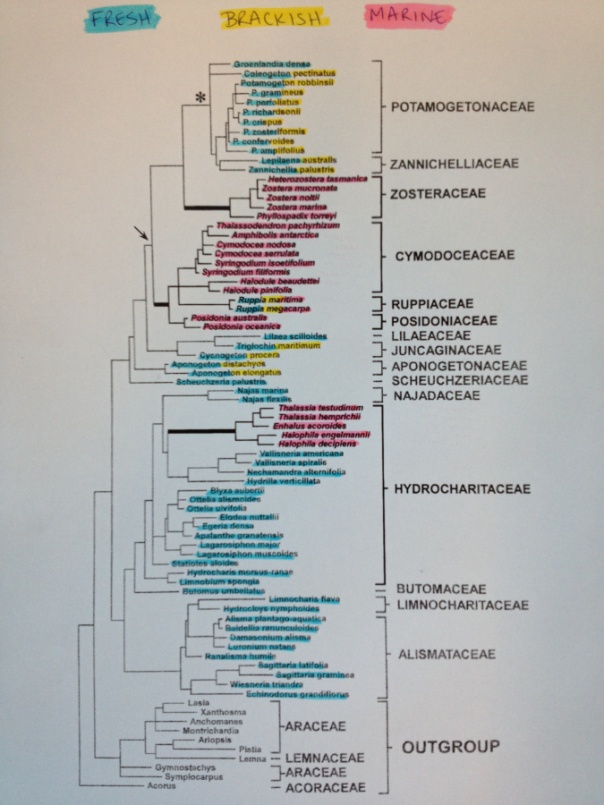
Plant-associated microorganisms are essential for their hosts’ survival and performance. Yet, most plant microbiome studies to date have focused on terrestrial species sampled across relatively small spatial scales. Here we report results of a global-scale analysis of microbial communities associated with leaf and root surfaces of the marine eelgrass Zostera marina throughout its range in the Northern Hemisphere. By contrasting host microbiomes with those of surrounding seawater and sediment, we uncovered the structure, composition and variability of microbial communities associated with eelgrass. We also investigated hypotheses about the assembly of the eelgrass microbiome using a metabolic modeling approach. Our results reveal leaf communities displaying high variability and spatial turnover, that mirror their adjacent coastal seawater microbiomes. In contrast, roots showed relatively low compositional turnover and were distinct from surrounding sediment communities — a result driven by the enrichment of predicted sulfur-oxidizing bacterial taxa on root surfaces. Predictions from metabolic modeling of enriched taxa were consistent with a habitat filtering community assembly mechanism whereby similarity in resource use drives taxonomic co-occurrence patterns on belowground, but not aboveground, host tissues. Our work provides evidence for a core eelgrass root microbiome with putative functional roles and highlights potentially disparate processes influencing microbial community assembly on different plant compartments.
IMPORTANCE Plants depend critically on their associated microbiome, yet the structure of microbial communities found on marine plants remains poorly understood in comparison to terrestrial species. Seagrasses are the only flowering plants that live entirely in marine environments. The return of terrestrial seagrass ancestors to oceans is among the most extreme habitat shifts documented in plants, making them an ideal test bed for the study of microbial symbioses with plants that experience relatively harsh abiotic conditions. In this study, we report results of a global sampling effort to extensively characterize the structure of microbial communities associated with the widespread seagrass species, Zostera marina or eelgrass, across its geographic range. Our results reveal major differences in the structure and composition of above- versus belowground microbial communities on eelgrass surfaces, as well as their relationships with the environment and host.