I gave at talk at SIO November 22, 2019 where I talked a bunch about seagrass microbiomes. I posted my slides to Slideshare and you can see them below.
Category Archives: Uncategorized
Investigating the Seagrass Microbiome
My blog post Investigating the Seagrass Microbiome, is a short summary of Cassie Ettinger’s 2017 paper on Zostera marina microbial communities. The post is a quick read that is written for a general audience and focuses on the paper’s main findings. I originally wrote the piece for my writing course (UWP 102B: Writing for Biological Sciences) and modeled it after the posts on the UCD Egghead Blog.
The post is published on the Egghead Blog and can be found here.
Slides for recent talk on our Seagrass Microbiome Project
Gave a talk recently trying to give an overview of this project.
I posted slides to Slideshare.
Here they are
And I also recorded the slides in synch with Audio using Camtasia. See
Preprint available: Microbiome succession during ammonification in eelgrass bed sediments
https://peerj.com/preprints/2956
Abstract
Background. Eelgrass (Zostera marina) is a marine angiosperm and foundation species that plays an important ecological role in primary production, food web support, and elemental cycling in coastal ecosystems. As with other plants, the microbial communities living in, on, and near eelgrass are thought to be intimately connected to the ecology and biology of eelgrass. Here we characterized the microbial communities in eelgrass sediments throughout an experiment to quantify the rate of ammonification, the first step in early remineralization of organic matter, or diagenesis, from plots at a field site in Bodega Bay, CA.
Methods. Sediment was collected from 72 plots from a 15 month long field experiment in which eelgrass genotypic richness and relatedness were manipulated. In the laboratory, we placed sediment samples (n= 4 per plot) under a N2 atmosphere, incubated them at in situ temperatures (15 oC) and sampled them initially and after 4, 7, 13, and 19 days to determine the ammonification rate. Comparative microbiome analysis using high throughput sequencing of 16S rRNA genes was performed on sediment samples taken initially and at 7, 13 and 19 days to characterize the relative abundances of microbial taxa and how they changed throughout early diagenesis.
Results. Within-sample diversity of the sediment microbial communities across all plots decreased after the initial timepoint using both richness based (observed number of OTUs, Chao1) and richness and evenness based diversity metrics (Shannon, Inverse Simpson). Additionally, microbial community composition changed across the different timepoints. Many of the observed changes in relative abundance of taxonomic groups between timepoints appeared driven by sulfur cycling with observed decreases in sulfur reducers (Desulfobacterales) and corresponding increases in sulfide oxidizers (Alteromonadales and Thiotrichales). None of these changes in composition or richness were associated with ammonification rates.
Discussion. Overall, our results showed that the microbiome of sediment from different plots followed similar successional patterns, which we surmise to be due to changes related to sulfur metabolism. These large changes likely overwhelmed any potential changes in sediment microbiome related to ammonification rate. We found no relationship between eelgrass presence or genetic composition and the microbiome. This was likely due to our sampling of bulk sediments to measure ammonification rates rather than sampling microbes in sediment directly in contact with the plants and suggests that eelgrass influence on the sediment microbiome may be limited in spatial extent. More in-depth functional studies associated with eelgrass microbiome will be required in order to fully understand the implications of these microbial communities in broader host-plant and ecosystem functions (e.g. elemental cycling and eelgrass-microbe interactions).
Now out in PeerJ: Microbial communities in sediment from Zostera marina patches, but not the Z. marina leaf or root microbiomes, vary in relation to distance from patch edge
https://peerj.com/articles/3246/?td=bl
tl;dr – The microbes (bacteria) on plant parts (root, leaf) and near-by sediment were different from each other. We did not find a difference between the microbes on eelgrass leaves or roots at the edge of a patch versus the middle of the patch. However, the microbes in sediments from different locations in the patch (middle, edge, outside of the patch) differed and these differences correlated with eelgrass density.
Abstract
Background
Zostera marina (also known as eelgrass) is a foundation species in coastal and marine ecosystems worldwide and is a model for studies of seagrasses (a paraphyletic group in the order Alismatales) that include all the known fully submerged marine angiosperms. In recent years, there has been a growing appreciation of the potential importance of the microbial communities (i.e., microbiomes) associated with various plant species. Here we report a study of variation in Z. marina microbiomes from a field site in Bodega Bay, CA.
Methods
We characterized and then compared the microbial communities of root, leaf and sediment samples (using 16S ribosomal RNA gene PCR and sequencing) and associated environmental parameters from the inside, edge and outside of a single subtidal Z. marina patch. Multiple comparative approaches were used to examine associations between microbiome features (e.g., diversity, taxonomic composition) and environmental parameters and to compare sample types and sites.
Results
Microbial communities differed significantly between sample types (root, leaf and sediment) and in sediments from different sites (inside, edge, outside). Carbon:Nitrogen ratio and eelgrass density were both significantly correlated to sediment community composition. Enrichment of certain taxonomic groups in each sample type was detected and analyzed in regard to possible functional implications (especially regarding sulfur metabolism).
Discussion
Our results are mostly consistent with prior work on seagrass associated microbiomes with a few differences and additional findings. From a functional point of view, the most significant finding is that many of the taxa that differ significantly between sample types and sites are closely related to ones commonly associated with various aspects of sulfur and nitrogen metabolism. Though not a traditional model organism, we believe that Z. marina can become a model for studies of marine plant-microbiome interactions.

Seagrass Microbiome Sampling
Recently the Seagrass Microbiome group has been wrapped up in sending (and receiving!) microbiome sampling kits. These kits are part of a larger collaborative project focused on re-sequencing of Zostera marina samples in conjunction with sequencing of additional marine and freshwater Alismatid species and their microbiomes. JGI recently sequenced and released the Zostera marina genome, and we are hoping to build on their efforts and explore population level variation within Zostera marina, as well as differences in genome content and structure between Zostera and other Alismatids, in conjunction with microbiome sequencing.
The sampling kits sent by the seagrass microbiome group have focused on the microbial aspect of this project. We have asked members of the Zostera Experimental Network (ZEN) as well as additional collaborators to sample both plant tissue for sequencing (coordinated through Jay Stachowicz and Jeanine Olsen) and microbiome samples. We are extremely excited about this sample set, as it covers populations of Zostera marina across many different environments, for which we already have extensive metadata through the ZEN group! We are requesting root, sediment (within the rhizosphere), and leaf tissue, as detailed in the diagram below (courtesy of Jeanine Olsen).

Collaborators are also sampling at two depths per site (deep and shallow), so that we can examine microbiome differences that may correlate with population depth. We are sampling 24 individuals per site, 12 per depth.
The kits are relatively straightforward and simple to both make and use, even if you’re not an experienced field microbiologist. We followed the kit and samplingl details we previously used (https://seagrassmicrobiome.org/protocols/microbial-sampling-kit/), with a few updates.
The kits now contain:
– 1 5cc syringe (for sediment collections)
– Tubes filled with Zymo buffer (DNA/RNA Shield)
– Plastic forceps
– Plastic spatula
– Parafilm
– Ethanol wipes
Here are a few photos of kit production:




We have sent out all of the kits, and have already started receiving some completed samples in the mail. Here is a close up of some of the samples from Kotzebue, AK.

A huge thanks to our collaborators for sampling, and to everyone from the Eisen lab who has helped make and send kits. Stay tuned for updates on sample processing and data !
Marine Algal and Plant Microbiomes Workshop – soliciting comments
So – am participating in a workshop, supported by CIFAR and the Gordon and Betty Moore Foundation over the next few days on “Marine Algal and Plant Microbiomes”. The workshop is basically trying to come up with a white paper / position paper on the future of such studies and to continue the conversation about this topic afterwards. We are asking questions like
- What are the challenges and opportunities in this area?
- What are the major scientific questions?
- How are such systems different from fresh water or terrestrial systems?
- How are they different?
- How are marine systems involving other hosts (e.g., coral, sponges, dolphins) comparable (i.e., is there something about marine systems that links them together in any way).
- What tools and resources could help advance work in this area?
And more
So I am posting here asking for a few bits of information from any readers.
- Are you interested in participating in follow up discussions on this topic?
- Do you know of any people we should try to bring into the conversation even if they are not, well, you?
- Are there any major projects in this area that would be worth engaging?
Any thoughts (on the topic that is) would be welcome.
Culturing Bacterial Isolates from the Seagrass Microbiome
My name is Karley Lujan and I am an undergraduate working on culturing bacterial isolates from the Seagrass microbiome. I joined this project because I am interested in learning about what information we can obtain from studying microbiomes. I think it is fascinating that although we can’t see microorganisms they are extremely prevalent and can have crucial roles in biological systems. The focus at the beginning of this project was to take Seagrass samples from Bodega Bay, create culture samples, and use Sanger sequencing of the 16S rRNA to identify what we grew. Seagrass and sediment samples were taken from Bodega Bay, CA. Then, in order to obtain isolates from the seagrass, we focused on the leaves, roots, and sediment. What we were able to successfully extract DNA from were identified as Shewanella, Pseudoalteromonas, Colwellia, Tenacibaculum, Vibrio and Alteromonas.


Methods:
- Sample preparation: Dilutions of sediment with PBS, PBS rinse of roots and leaves, ground and crushed leaves with PBS
- Culturing: Plated the PBS sample solutions onto two of each of the following plate types; one plate for 25℃ and the other at 4℃
Agar Plates/Liquid Media
- Nitrogen Free
- Seawater Agar
- Seawater Medium
- Difco Marine Broth
- Selected Colonies: After there was significant growth on the plates we selected various interesting colonies and isolated them by dilution streaking. Single colonies were then grown overnight in the appropriate liquid media and at the appropriate temperature
- DNA Extraction: Genomic DNA extractions were performed and glycerol stocks were made using the successful liquid cultures. Extracted DNA then went through 16S rRNA gene PCR and gel electrophoresis in order to confirm that enough DNA was present for Sanger Sequencing
- Sanger Sequencing: 16S rRNA sequences for each isolate were ran through BLAST and phylogenetic trees were built in order to obtain tentative identifications for the isolates
Results: After Sanger sequencing, the data was ran through BLAST to obtain a tentative identification and determine whether or not the microbe was a good candidate for sequencing.
Shewanella: Electrogenic- An electron generator that can be used in microbial fuel cells.
Colwellia: Cold-adapted
Vibrio: Some species of Vibrio can go through morphogenetic changes after going from a liquid to a solid surface. This leads them to change from swimmer cells to swarmer cells.
Pseudomonas: Two bacterial isolates were cable of growing on Nitrogen-Free agar plates at 25⁰C. Identified as part of the genera Pseudomonas, there are some species of this genera capable of aerobically fixing nitrogen. These are of particular interest as we will be further investigating which nitrogen-fixing bacteria are essential for seagrass health.
Currently I am beginning to look at the genomes of the bacteria we decided to sequence and I am also working with bacteria that are capable of growing on the nitrogen-free agar plates. At first it was difficult to extract the DNA from these bacteria but now both have been tentatively identified as Pseudomonas through sanger sequencing of the 16S gene. This is interesting because there aren’t many Pseudomonas that can fix nitrogen which is what these two must be doing in order to survive on the nitrogen-free plates. These two bacteria also have different morphologies which means they could be different species in the genus Pseudomonas. Due to their morphological similarities yet ability to grow on nitrogen-free agar, I think these two bacteria are very interesting and we will be finding out more about them by sequencing and analyzing their genomes.
CA sampling for project phylogeny
A few of us at Team Seagrass have been doing fieldwork in Northern CA in order to collect seagrass relatives. We have specifically been targeting freshwater species (highlighted in blue in the below phylogeny). We will continue with marine and brackish species when the tides in the San Francisco Bay are lower.
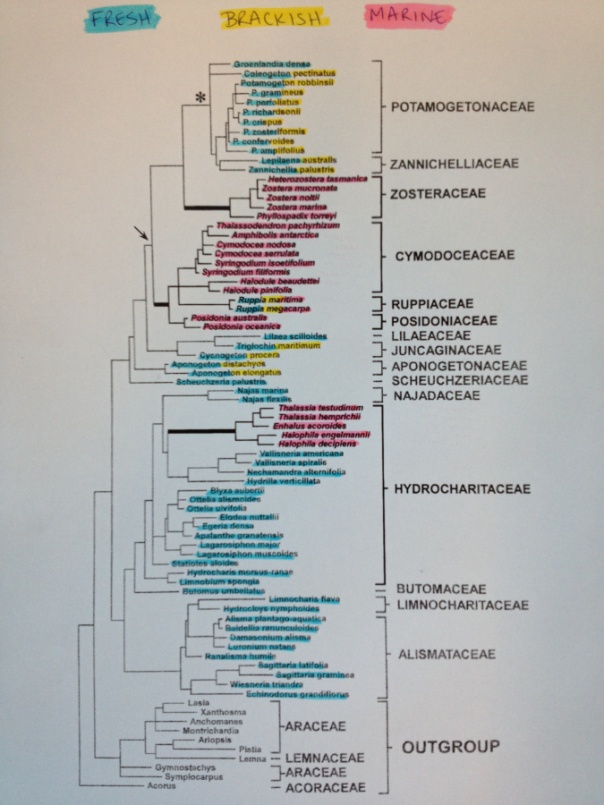
We have been following the methods detailed in https://seagrassmicrobiome.org/sample-collection-and-preservation/ and have managed to collect a few representative species from the following clades: Potamogetonaceae, Alismataceae, Hydrocharitaceae, Najadaceae, and the Lilaceae.

For further plant identification and the construction of a ‘host phylogeny’ we will be using chloroplast DNA markers.
In addition to what we have sampled thus far, we have also managed to sample some outgroups (Myriophyllum, Ceratophyllum, and Camboba). We will hopefully be able to sample additional outgroup species shown on the above tree.
We have been successful in 7 out of the 10 locations that we have tried. The lack of success so far can be attributed to drought conditions in CA and the increase in invasive submerged aquatic vegetation (e.g. Myriophyllum, Water Milfoil; and Eichhornia crassipes, Water Hyacinth). Water Milfoil and Water Hyacinth form dense mats below and above water, respectively, outcompeting native vegetation. Even in sites where we been successful, much of the area we surveyed has been overrun with these two invasive species.
Obviously drought has also been an issue for us, as many of the lakes and tributaries have little to no water in them this year, resulting in a lack of any aquatic vegetation. Water levels in Folsom Lake, for example, are so far reduced that the Park Service has had to build additional parking lots on the water side of the boat ramp in order for people to be able to get close to the new shoreline.

In the following weeks we will head up to the foothills in search of remaining freshwater species, as well as explore the Bay Area coastline and salt marsh/Delta area for marine and brackish species. Stay tuned!
Sample preservation experiment
During ZEN DNA extractions we noticed that samples preserved in the Zymo buffer were forming a precipitate with the C1 solution from the MoBio kit. Furthermore, many of the samples also resulted in very low DNA yields, perhaps correlated with the precipitate formation. Any microbiologist will tell you that there are many different ways to preserve samples from the field, but there does not seem to be a universal *best* method.
We decided the best way to approach this problem for the Seagrass Microbiome project was to explore a variety of sample preservation methods and see which approximated the ‘real’ microbiome best (as measured by preservation on dry ice). We chose the following methods to try: Dry ice, Zymo, RNA-later, Drierite, and Ethanol. We also wanted to investigate how different preservation methods performed over time. We chose 4 time points post sampling for our extractions: 24 hours, 1 week, 2 weeks, and 1 month.
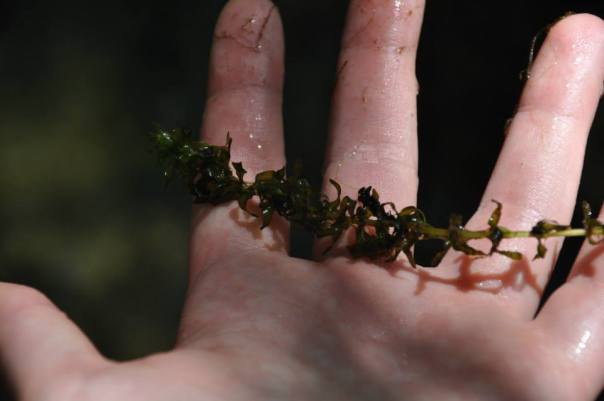
We drove to Putah Creek in Winters in search of submerged aquatic plants in the Alismatales (the order that contains the seagrasses). Here is a photo of our study site: 
We found a bed of Elodea canadensis growing near the shore and started sampling.
We pulled out whole plants and divided them into root and leaf for each of our trial preservation methods, with 3 replicates per method per time point. We extracted DNA at each of the different time points, and did PCR of the 16S SSU rRNA gene, with subsequent library preparation and sequencing on the Illumina Mi-Seq. We used QIIME to analyze the sequence data and compare species diversity between our preservation methods.
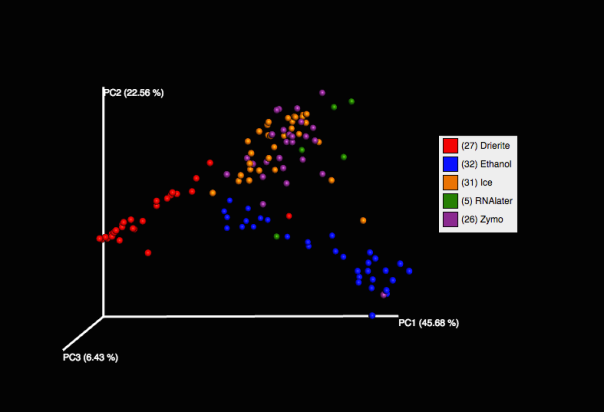
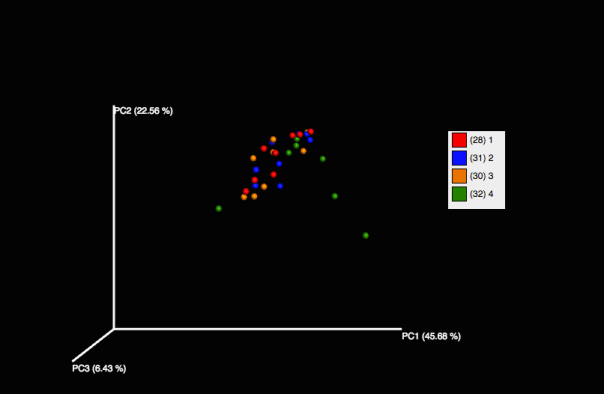
Our PCoA plot shows that preservation in Zymo best approximates the community captured using dry ice (our control), and that Drierite and Ethanol were both very different from anything else. RNA-later appears close to Zymo and dry ice, but we only have 5 data points for it. For the RNA-later and Ethanol extractions, we extracted directly from the solution without pelleting, which may have affected our end result.
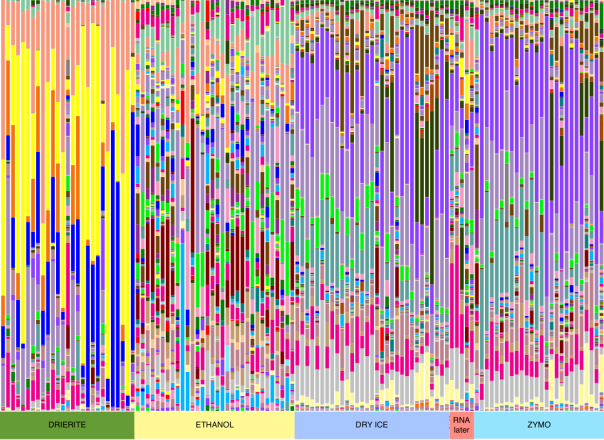
The taxonomy table supports the trend seen in our PCoA plot, with Drierite and Ethanol being very different, and Zymo, dry ice, and our 5 RNA-later time points showing similar communities.
Looking at variation in community structure across time points within Zymo we see little to no change.
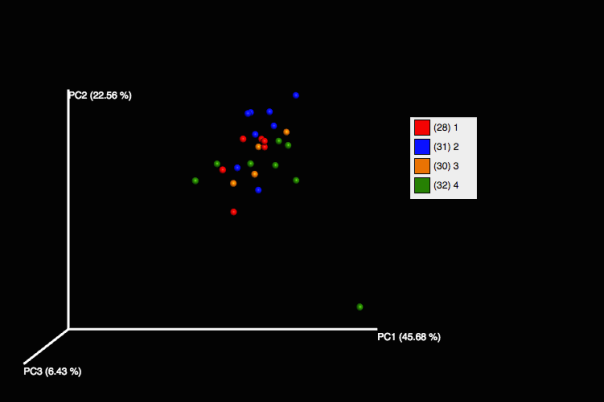
However, we see some change in community structure by the month mark with dry ice.
Take home messages from our sample preservation experiment:
- Using Drierite as a preservation technique does not capture the community assembly well.
- RNA-later and Ethanol could have had better success with pelleting and removing the supernatant prior to extraction. We may investigate this further in the future.
- In spite of our initial concerns regarding precipitate forming in the Zymo buffer, Zymo is the clear winner in our trial.
- There are slight changes in the community assembly by the month mark, in all methods (depending on whether you use weighted or unweighted unifrac PCoA).
- Due to the current ease of sampling using dry ice, we will continue this method except in situations where samples must be kept at room temperature (we will then use Zymo).
HAPPY SAMPLING 🙂 Please see https://seagrassmicrobiome.org/sample-collection-and-preservation/ for more details regarding sampling and preservation protocols.

